Introduction
Genetic variation describes naturally occurring genetic differences among individuals of the same species. All organisms (yup even ones that look really similar like sheep) are slightly or greatly different. This variation within a population is a biological advantage as it provides a buffer and ensures the survival of some individuals in the face of changing environmental circumstances. In New Zealand the habitat is constantly changing and living (biotic) and non living (abiotic factors) change the populations gene pools and provide selection pressures.
This standard examines what influences the variation in populations and how this leads to different frequencies of traits and eventually natural selection. What leads to the variation in a species?
Genetic variation describes naturally occurring genetic differences among individuals of the same species. All organisms (yup even ones that look really similar like sheep) are slightly or greatly different. This variation within a population is a biological advantage as it provides a buffer and ensures the survival of some individuals in the face of changing environmental circumstances. In New Zealand the habitat is constantly changing and living (biotic) and non living (abiotic factors) change the populations gene pools and provide selection pressures.
This standard examines what influences the variation in populations and how this leads to different frequencies of traits and eventually natural selection. What leads to the variation in a species?
Let's look at variation
The amount of variations affects the survivability of that population. Low genetic biodiversity means that during states of environmental change a population is less likely to survive.
The amount of variations affects the survivability of that population. Low genetic biodiversity means that during states of environmental change a population is less likely to survive.
Sources of variation within a gene pool
A gene pool is the complete set of unique alleles within a population. The genetic diversity is the measure of different alleles present in that gene pool. How does genetic variation increase or decrease within a gene pool? And what effect do fluctuations in genetic variation have on populations over time?
Genetic code Click on the button below to review the key ideas surrounding DNA and genetics:
A gene pool is the complete set of unique alleles within a population. The genetic diversity is the measure of different alleles present in that gene pool. How does genetic variation increase or decrease within a gene pool? And what effect do fluctuations in genetic variation have on populations over time?
Genetic code Click on the button below to review the key ideas surrounding DNA and genetics:
and revisit some terminology from year 11:

Remember the base sequence of DNA carries the instructions to make proteins. These proteins determine the growth, development and characteristics of an organism. A gene is a length of DNA which codes for a particular protein. Alternative forms of a gene which code for this protein are called alleles. The reason different forms of a gene are expressed differently for that single trait is because of the varied sequence of nucleotides within that gene.
Sexually reproducing individuals pass this genetic information in varying combinations via their gametes. Offspring are genetically different from their parents due the fact that during meiosis, random fertilisation of gametes, mutations and the environment.
If a mutation occurs in body cells they are a somatic mutation during the lifetime of the organism it will not be passed on to the next generation. If a mutation occurs in the sex cell it is a gametic mutations and can be passed onto offspring. This mutation will be incorporated into every cell of that offspring. REMEMBER neutral or silent mutations have no observable effect, beneficial mutations may provide an adaptive advantage to an organism so that the mutation is passed on through generations because it gives that individual a reproductive advantage! (more babies more often). However REMEMBER a mutation will ONLY be inherited by offspring if the gamete which contains that mutation fuses with another gamete. This is how variation is produced and it changes/increases the allele frequency in the gene pool. Harmful mutations which affect the survival of an organism will be selected against and may not be passed on therefore decreasing the frequency of that allele.
Check out this link HERE
Sexually reproducing individuals pass this genetic information in varying combinations via their gametes. Offspring are genetically different from their parents due the fact that during meiosis, random fertilisation of gametes, mutations and the environment.
- Mutations
If a mutation occurs in body cells they are a somatic mutation during the lifetime of the organism it will not be passed on to the next generation. If a mutation occurs in the sex cell it is a gametic mutations and can be passed onto offspring. This mutation will be incorporated into every cell of that offspring. REMEMBER neutral or silent mutations have no observable effect, beneficial mutations may provide an adaptive advantage to an organism so that the mutation is passed on through generations because it gives that individual a reproductive advantage! (more babies more often). However REMEMBER a mutation will ONLY be inherited by offspring if the gamete which contains that mutation fuses with another gamete. This is how variation is produced and it changes/increases the allele frequency in the gene pool. Harmful mutations which affect the survival of an organism will be selected against and may not be passed on therefore decreasing the frequency of that allele.
Check out this link HERE
Sexual Reproduction
The process of sexual reproduction involves two parents, each contributing one gamete. Gametes are produced by a process called meiosis, which starts by the duplication of the chromosomes, followed by two rounds of cell divisions and halving of the chromosome number. Gametes have haploid chromosome number. Meiosis produces genetic variation within a species.
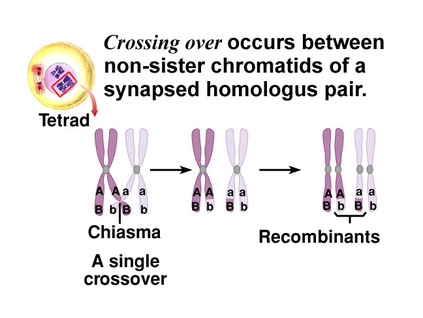
A key feature of meiosis is the exchange of chromosome segments between the closest homologous chromatids during first division of this process. This is called crossing over resulting in the recombination of DNA. Recombination shuffles the allele content between sister chromatids. BECAUSE crossing over is random, its effect on the level of variation contributed to gametes is different in each case. Here is a great animation: Here
Independent assortment is the major source of genetic variability in gametes which are inherited by offspring. It occurs when each of the homologous chromosome pairs up randomly each side of the equator meaning that when they separate (segregate) in the first division the combination of alleles which end up in each gamete are unique. The homologous pairs do this independently of each other creating a huge amount of variation in the nature of the gametes produced.
Eg, in humans there are 23 pairs of chromosomes, 8 million)
Check out this great animation HERE
The process of sexual reproduction involves two parents, each contributing one gamete. Gametes are produced by a process called meiosis, which starts by the duplication of the chromosomes, followed by two rounds of cell divisions and halving of the chromosome number. Gametes have haploid chromosome number. Meiosis produces genetic variation within a species.
A key feature of meiosis is the exchange of chromosome segments between the closest homologous chromatids during first division of this process. This is called crossing over resulting in the recombination of DNA. Recombination shuffles the allele content between sister chromatids. BECAUSE crossing over is random, its effect on the level of variation contributed to gametes is different in each case. Here is a great animation: Here
Independent assortment is the major source of genetic variability in gametes which are inherited by offspring. It occurs when each of the homologous chromosome pairs up randomly each side of the equator meaning that when they separate (segregate) in the first division the combination of alleles which end up in each gamete are unique. The homologous pairs do this independently of each other creating a huge amount of variation in the nature of the gametes produced.
Eg, in humans there are 23 pairs of chromosomes, 8 million)
Check out this great animation HERE
Year 12s go Cartooloony
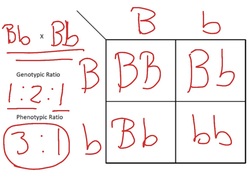
- Monohybrid inheritance is the inheritance of ONE characteristic/trait controlled by ONE gene at a particular locus on a homologous chromosome. The gene will have alternative forms called alleles. The alleles will be dominant or recessive. An organisms genotype is determined by the combination of alleles inherited from each parent.
Test Cross: Is an organism displays a dominant trait how do you know if they are homozygous dominant (pure breed) or heterozygous? DO A TEST CROSS OF COURSE!!
A test cross with a homologous recessive individual. You can be fairly certain of the phenotype of the parent (BB or Bb), HOWEVER due to the random nature of fertilisation offspring may not reflect predicted outcomes in the first generation of offspring.
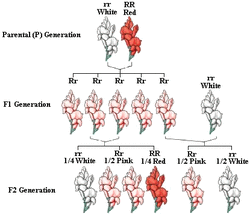
Incomplete dominance: is a form of intermediate inheritance in which one allele for a specific trait is not completely dominant over the other allele. This results in a third phenotype in which the expressed physical trait is a combination of the dominant and recessive phenotypes in the heterozygous genotype.
Note peeps: How do the genotype and phenotype ratios for heterozygous x heterozygous compare with that of the same cross between complete dominant alleles?
Co-dominance: occurs when both alleles are equally dominant and both fully expressed if present in a heterozygous genotype. This results in a third phenotype in which the exhibit both physical trait.
Multiple alleles: This is super interesting when genes have more than two different forms of a gene (alleles).
Blood typing is a great example of increased variation in the human race:
AB blood is the codominant relationship between the A protein and B protein both expressing themselves completely.
AO (type O allele means there is no protein), A is dominant and you see type A phenotype.
BO is the same except you see the B phenotype. Type O is recessive.
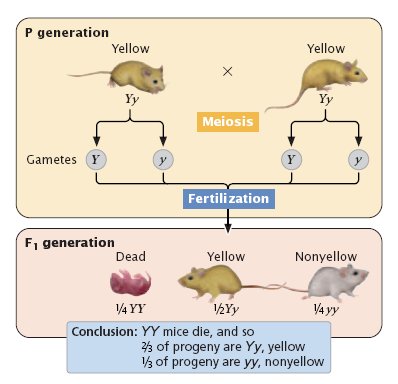
Lethal alleles: occur when a mutation results in an allele that produces a non functional version of an essential protein. If the individual inherits a lethal combination of this allele it will die shortly after birth/germination.
Note peeps: How do the genotype and phenotype ratios for heterozygous x heterozygous compare with that of the same cross between complete dominant alleles?
Co-dominance: occurs when both alleles are equally dominant and both fully expressed if present in a heterozygous genotype. This results in a third phenotype in which the exhibit both physical trait.
Multiple alleles: This is super interesting when genes have more than two different forms of a gene (alleles).
Blood typing is a great example of increased variation in the human race:
AB blood is the codominant relationship between the A protein and B protein both expressing themselves completely.
AO (type O allele means there is no protein), A is dominant and you see type A phenotype.
BO is the same except you see the B phenotype. Type O is recessive.
Lethal alleles: occur when a mutation results in an allele that produces a non functional version of an essential protein. If the individual inherits a lethal combination of this allele it will die shortly after birth/germination.
Individuals living in populations share a gene pool will exhibit genetic biodiversity. Genetic biodiversity is the measure of the different genetic combinations within the gene pool. The amount of genetic biodiversity (variation) within a population influences the likelihood of survival of that population.
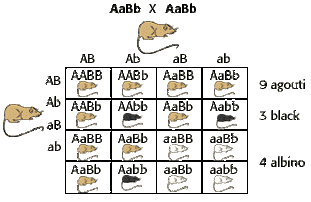
Dihybrid Inheritance
Dihybrid crosses are genetic crosses involving the inheritance of two genes and, therefore, two traits. Dihybrid crosses show how the production of these gametes bring about the inheritance of traits. Four types of gamete are produced for each parent. This is the result of independent assortment of the homologous pairs of chromosomes and their separation during the process of meiosis.
Linked Genes:
The dihybrid cross we previously did assumed the genes were on different pairs of chromosomes. Now, we want to look at an example where the genes involved are on the same chromosome. One such example is the flower color and pollen shape experiment done by Bateson and Punnett. In the plants that they studied, the genes for pollen shape and flower color are located on the same chromosome (pair) as each other, thus are inherited together.
Linked Genes If the parents are PPLL × ppll, the first parent will only make gametes with PL and the second with pl, which doesn’t seem too different so far. From these parents, the F1 generation would all be PpLl. However, when calculating what the F2 generation will be, since the genes are located on the same pair of chromosomes, then theoretically, the only possible gametes are PL and pl (not Pl or pL). So
PL pl
PL PPLL PpLl
pl PpLl ppll
The phenotype ratio for this cross is 3:1, not 9:3:3:1 as would be expected for a “normal” dihybrid cross. Because these genes are on the same chromosome pair, they are called linked genes. Interestingly, Bateson and Punnett’s results showed just a few, unexpected ppL- and P-ll offspring, more than would be predicted by linked genes, but far less than would be predicted by unlinked genes in a “regular” dihybrid cross. This is due to the fact that occasionally, during synapsis in meiosis I, while the homologous chromosomes are paired up, sister chromatids from the homologous chromosomes exchange equal segments. This is called crossing over. In the flower example, a few of the plants could exhibit crossing over during meiosis I, producing a few pL and Pl gametes, which would account for the small number of ppL- and P-ll offspring. T. H. Morgan and his grad students, who studied fruitflies, found that the farther apart two genes are on a chromosome, the more likely there is to be crossing over between those two genes. They found that for any given two genes on the same chromosome as each other, the amount of crossing over that occurs is a fairly constant quantity that can be measured. From their crossing over data, Morgan et al. were able to arrange fruit fly genes in the order in which they occur on the fruit fly chromosomes. Interestingly, if two genes are very far apart on the same chromosome pair, there is so much crossing over that the results obtained look like a regular dihybrid cross between unlinked genes.
THINGS TO REMEMBER:
Linked genes / alleles are on the same chromosome.
They do not assort independently (like genes on different chromosomes, therefore ratio is not 9:3:3:1) as they are physically linked to each other on the same chromosome and cannot separate (randomly) during meiosis.
Crossing over exchanges genetic information of the chromosome between the two chromatids during meiosis when homologous chromosomes are paired up. Crossing over produces the Pl and pL gametes / new combination of alleles OR well-annotated diagram.
The low occurrence indicates the genes / alleles must be close together on the same chromosome because the probability of crossing over / number of recombinants increases as gene distance increases.
Dihybrid crosses are genetic crosses involving the inheritance of two genes and, therefore, two traits. Dihybrid crosses show how the production of these gametes bring about the inheritance of traits. Four types of gamete are produced for each parent. This is the result of independent assortment of the homologous pairs of chromosomes and their separation during the process of meiosis.
Linked Genes:
The dihybrid cross we previously did assumed the genes were on different pairs of chromosomes. Now, we want to look at an example where the genes involved are on the same chromosome. One such example is the flower color and pollen shape experiment done by Bateson and Punnett. In the plants that they studied, the genes for pollen shape and flower color are located on the same chromosome (pair) as each other, thus are inherited together.
Linked Genes If the parents are PPLL × ppll, the first parent will only make gametes with PL and the second with pl, which doesn’t seem too different so far. From these parents, the F1 generation would all be PpLl. However, when calculating what the F2 generation will be, since the genes are located on the same pair of chromosomes, then theoretically, the only possible gametes are PL and pl (not Pl or pL). So
PL pl
PL PPLL PpLl
pl PpLl ppll
The phenotype ratio for this cross is 3:1, not 9:3:3:1 as would be expected for a “normal” dihybrid cross. Because these genes are on the same chromosome pair, they are called linked genes. Interestingly, Bateson and Punnett’s results showed just a few, unexpected ppL- and P-ll offspring, more than would be predicted by linked genes, but far less than would be predicted by unlinked genes in a “regular” dihybrid cross. This is due to the fact that occasionally, during synapsis in meiosis I, while the homologous chromosomes are paired up, sister chromatids from the homologous chromosomes exchange equal segments. This is called crossing over. In the flower example, a few of the plants could exhibit crossing over during meiosis I, producing a few pL and Pl gametes, which would account for the small number of ppL- and P-ll offspring. T. H. Morgan and his grad students, who studied fruitflies, found that the farther apart two genes are on a chromosome, the more likely there is to be crossing over between those two genes. They found that for any given two genes on the same chromosome as each other, the amount of crossing over that occurs is a fairly constant quantity that can be measured. From their crossing over data, Morgan et al. were able to arrange fruit fly genes in the order in which they occur on the fruit fly chromosomes. Interestingly, if two genes are very far apart on the same chromosome pair, there is so much crossing over that the results obtained look like a regular dihybrid cross between unlinked genes.
THINGS TO REMEMBER:
Linked genes / alleles are on the same chromosome.
They do not assort independently (like genes on different chromosomes, therefore ratio is not 9:3:3:1) as they are physically linked to each other on the same chromosome and cannot separate (randomly) during meiosis.
Crossing over exchanges genetic information of the chromosome between the two chromatids during meiosis when homologous chromosomes are paired up. Crossing over produces the Pl and pL gametes / new combination of alleles OR well-annotated diagram.
The low occurrence indicates the genes / alleles must be close together on the same chromosome because the probability of crossing over / number of recombinants increases as gene distance increases.
Here is a great little animation: HERE
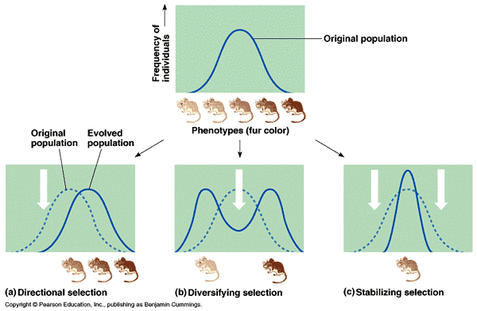
The overall effect
How do populations respond to all these forces? As relative allele frequencies change, relative genotype frequencies may also change. Each genotype in the population usually has a different fitness for that particular environment. In other words, some genotypes will be favored, and individuals with those genotypes will continue to reproduce and these advantageous alleles will increase within the population. Other genotypes will not be favored: individuals with those genotypes will be less likely to reproduce or die and there fore reduce the number of those alleles within a population. This is natural selection. Here is a little animation to view: Here
What type of genotype would be unfavorable? Unfavorable genotypes take many forms, such as increased risk of predation, decreased access to mates, or decreased access to resources that maintain health. Overall, the forces that cause relative allele frequencies to change at the population level can also influence the selection forces that shape them over successive generations.
For example, if moths with genotype aa migrate into a population composed of AA and Aa individuals, they will increase the relative allele frequency of a. However, if the aa genotype has a clear disadvantage to survival (e.g. vulnerability to predation), eventually the changes brought about by the initial migration will be reversed.
Types of natural selection include:
How do populations respond to all these forces? As relative allele frequencies change, relative genotype frequencies may also change. Each genotype in the population usually has a different fitness for that particular environment. In other words, some genotypes will be favored, and individuals with those genotypes will continue to reproduce and these advantageous alleles will increase within the population. Other genotypes will not be favored: individuals with those genotypes will be less likely to reproduce or die and there fore reduce the number of those alleles within a population. This is natural selection. Here is a little animation to view: Here
What type of genotype would be unfavorable? Unfavorable genotypes take many forms, such as increased risk of predation, decreased access to mates, or decreased access to resources that maintain health. Overall, the forces that cause relative allele frequencies to change at the population level can also influence the selection forces that shape them over successive generations.
For example, if moths with genotype aa migrate into a population composed of AA and Aa individuals, they will increase the relative allele frequency of a. However, if the aa genotype has a clear disadvantage to survival (e.g. vulnerability to predation), eventually the changes brought about by the initial migration will be reversed.
Types of natural selection include:
|
|
|
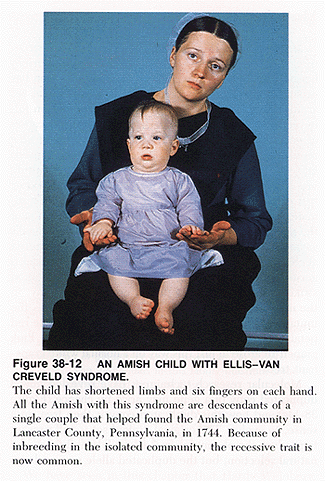
Factors which will affect allele frequencies within a gene pool:
Typically, genetic drift occurs in small populations, where infrequently-occurring alleles face a greater chance of being lost. Once it begins, genetic drift will continue until the involved allele is either lost by a population or is the only allele present at a particular gene locus within a population. Both possibilities decrease the genetic diversity of a population.
Genetic drift is common after a population experiences a population bottleneck. A population bottleneck arises when a significant number of individuals in a population die or are otherwise prevented from breeding, resulting in a drastic decrease in the size of the population. Genetic drift can result in the loss of rare alleles, and can decrease the size of the gene pool.
- Migration
- Random forces lead to genetic drift
Typically, genetic drift occurs in small populations, where infrequently-occurring alleles face a greater chance of being lost. Once it begins, genetic drift will continue until the involved allele is either lost by a population or is the only allele present at a particular gene locus within a population. Both possibilities decrease the genetic diversity of a population.
Genetic drift is common after a population experiences a population bottleneck. A population bottleneck arises when a significant number of individuals in a population die or are otherwise prevented from breeding, resulting in a drastic decrease in the size of the population. Genetic drift can result in the loss of rare alleles, and can decrease the size of the gene pool.